Development of gel-based multiplex RT-PCR for detection of ER/PR/HER2-positive breast cancer for diagnosis
Suh Kyung (Stephanie) Yoon
Hong Kong International School
1 Red Hill Road, Tai Tam, Hong Kong
[email protected]
Abstract
Estrogen receptor (ER), Progesterone receptor (PR), and Human epidermal growth factor receptor 2 (HER2) expression level plays a central role as a prognostic and predictive marker in breast cancer specimens. Therefore, detecting ER, PR, and HER2 status is essential for determining a correct therapeutic method to treat breast cancer. The most commonly used assays in the clinical studies for detecting HER2 status are immunohistochemistry (IHC) and fluorescence in situ hybridization (FISH). However, there are controversial issues involving cases with ER, PR, and HER2 status based on these semi-quantitative methods. IHC and FISH frequently underestimate ER, PR, and HER2 status. We developed a multiplex quantitative and cost-effective analysis of gel-based reverse transcription-polymerase chain reaction (RT-PCR) for the assessment of ER, PR, and HER2 in four breast cancer cell lines. Multiplex RT-PCR provided consistent data in breast cancer cell lines without any cross-amplification of cDNA on other genes indicating that our developed assay was reliable in specificity. In fact, this multiplex RT-PCR assay proved to be a sensitive and convenient method to rapidly and simultaneously detect four genogroups of human ER, PR, HER2, and Pumilio homolog 1 (PUM1), which is known to be a normalizing gene in breast cancer. In conclusion, multiplex RT-PCR could be certainly useful for routine diagnosis of ER, PR, and HER2 positive breast cancer.
Introduction
The advancement and widespread application of genomics, transcriptomics, and proteomics has provided novel understanding of the molecular complexity of breast cancer. However, clinical decisions still rely on assessment of three molecular markers: estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor 2 (HER2) [1]. Identifying these markers is essential for efficient targeted treatment of the different types of breast cancer.
ER is a receptor that is activated by estrogen. There are two types of ER: ERα and ERβ. Once activated, ERα and ERβ form dimers and translocate into the nucleus to facilitate the regulation of various genes. Approximately 80% of breast cancer patients display ER-positive breast cancer. ER-positive breast cancer typically responds much more readily to hormone-targeted therapy. It is treated with tamoxifen, which is a drug that blocks hormone receptors, causing the inhibition of estrogen production [2].
PR is a receptor that blocks transcription until activated by progesterone. Approximately 65% of breast cancer patients display PR positive breast cancer. While PR may not respond to endocrine therapy directly, their activation may have significant impacts on the ER signaling pathway, thus showing value in their determination of which tumors may be subject to PR reprogramming of ER [2].
HER2 is a breast cancer diagnostic marker that aids the therapeutic decisions in the treatment of breast cancer [3]. Approximately 10-30% of patients display HER2 positive breast cancer. HER2 gene amplification in breast cancer cells is linked to a more clinically aggressive response in patients and corresponds to a higher death rate [4]. HER2 positive breast cancer is normally treated with trastuzumab, a monoclonal antibody that induces the downregulation and internalization of HER2, as well as upregulates cell cycle inhibitors [4].
Previously, detection of ER, PR, and HER2 in breast cancer cells has been widely done using the Immunohistochemistry (IHC) and Fluorescence in situ hybridization (FISH) methods. However, immunostaining methods such as IHC are naturally prone to error as the results are semi-quantitative and subject to interobserver variability [5]. Thus, these methods often abuse of neglect patients. FISH assessments give much more sensitive and quantitative results, but are expensive and time-consuming, thus not appropriate for widespread use. Furthermore, FISH methods are also prone to ambiguous results as standard protocols for sample collection and storage have not been developed yet.
Clearly, detection of receptor DNA or cDNA by PCR meets the criteria of speed and high sensitivity, so it is used frequently for diagnostic purposes. Gel-based reverse transcription polymerase chain reaction (RT-PCR) assay was reported for the diagnosis of African swine fever (ASF) [6,7]. This assay was sensitive and specific for the fast and early diagnosis of the ASF. Therefore, RT-PCR could be an alternative method for the detection of ER, PR, and HER2, as it is an inexpensive test that yields sensitive and accurate results [8]. Furthermore, multiplex PCR allows for simultaneous amplification of multiple target sequences in a single tube using specific primer sets in combination. Thus, multiplex PCR favors higher throughput and automation compared to singleplex PCR, especially when testing repeated and numerous patient analyses of the same targets [9]. To overcome the problems of two-step real-time singleplex RT-PCR, a highly sensitive and specific gel-based multiplex RT-PCR assay was developed in this study by testing ER, PR, and HER2 negative and positive breast cancer cell lines.
The Goal of the Project
Method and Experiments
Breast cancer cell lines
4 different known breast cancer cell lines were used in this study as templates for the HER2 detection, being the MCF7, MDAMB231, BT474, and MDAMB453. These cell lines all display different types of cell markers (ER, PR, HER2), being HER2-, triple negative, triple positive, and HER2+ respectively (Table 1).
Primer set design
Primers were developed for ER, PR, HER2 and PUM1 (Table 2). They were chosen as all different lengths so they could be differentiated during agarose gel electrophoresis, with ER - 244 bp, PR - 224, HER2 - 342, and PUM1 - 187. Their annealing temperatures were fairly similar, all ranging around 60°C.
RT-PCR reaction
RNA was extracted from breast cancer cell lines by RNA-spin™ Total RNA Extraction Kit (Intron) following the manufacturer's instructions. cDNA was synthesized from the extracted RNA using TOPscript™ Reverse Transcriptase (Enzynomics) following the manufacturer’s instructions. For the preparation for the PCR reaction, the different primers were tested individually first using 20 µL reaction containing 2µL forward/reverse primers (10 pmol), 2µL reaction buffer, 2µL dNTP, 1.125µL Taq polymerase(Bioneer), 0.5 µL cDNA, and RNase Free dH2O up to 20 µL. PCR was done with annealing temperature 60°C and an extension time of 40 seconds in 72°C for 40 cycles. For the multiplex reactions, nTaq-multi HOT (Enzynomics) was used with 20 µL reaction containing 2µL forward/reverse primers, 0.2µL nTaq- multiHOT polymerase, 0.5 µL cDNA, and RNase Free dH2O up to 20 µL.
Agarose gel electrophoresis
2-5% agarose gel and TBE buffer were used in this study. RedSafe™ Nucleic Acid Staining Solution (Intron), which is an alternative to the traditional ethidium bromide(EtBr), was used to stain nucleic acid. Amplified DNA was detected by direct examination of the gel in ultraviolet(UV) light.
Agarose Gel Quantification Analysis
LI-COR Image Studio software version 2.1.10 and gel images were saved as a work area. All gel intensity quantification analyses were performed on images saved in TIFF format.
All experiments in this study including running the agarose gel electrophoresis were performed in University of Suwon, Korea.
Results
With the molecular classifications of breast cancer, researchers focus on breast cancer cell lines to determine whether the molecular profiles observed in breast cancer patients are reflected in cell line models of the disease. Applications of transcriptional profiling to breast cancer cell lines using various platforms provided the characteristics of the cell lines by the expression of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2). Therefore, 4 cell lines were selected to observe ER, PR, and HER2 expressions with RT-PCR method (Table 1).
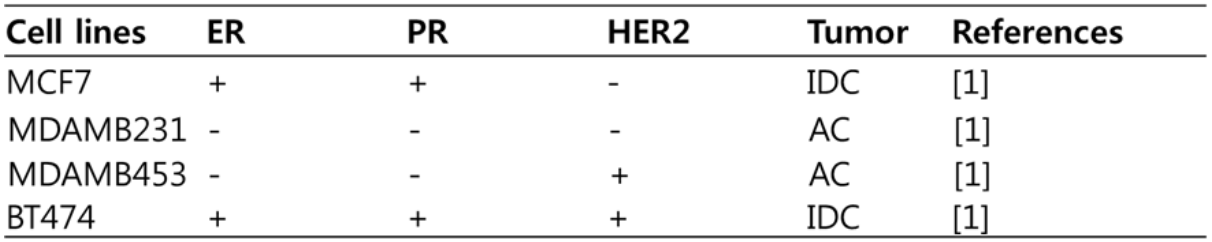
Table 1. Categorization and molecular information of 4 breast cancer cell lines, and the clinical features of tumors where they derive
Hong Kong International School
1 Red Hill Road, Tai Tam, Hong Kong
[email protected]
Abstract
Estrogen receptor (ER), Progesterone receptor (PR), and Human epidermal growth factor receptor 2 (HER2) expression level plays a central role as a prognostic and predictive marker in breast cancer specimens. Therefore, detecting ER, PR, and HER2 status is essential for determining a correct therapeutic method to treat breast cancer. The most commonly used assays in the clinical studies for detecting HER2 status are immunohistochemistry (IHC) and fluorescence in situ hybridization (FISH). However, there are controversial issues involving cases with ER, PR, and HER2 status based on these semi-quantitative methods. IHC and FISH frequently underestimate ER, PR, and HER2 status. We developed a multiplex quantitative and cost-effective analysis of gel-based reverse transcription-polymerase chain reaction (RT-PCR) for the assessment of ER, PR, and HER2 in four breast cancer cell lines. Multiplex RT-PCR provided consistent data in breast cancer cell lines without any cross-amplification of cDNA on other genes indicating that our developed assay was reliable in specificity. In fact, this multiplex RT-PCR assay proved to be a sensitive and convenient method to rapidly and simultaneously detect four genogroups of human ER, PR, HER2, and Pumilio homolog 1 (PUM1), which is known to be a normalizing gene in breast cancer. In conclusion, multiplex RT-PCR could be certainly useful for routine diagnosis of ER, PR, and HER2 positive breast cancer.
Introduction
The advancement and widespread application of genomics, transcriptomics, and proteomics has provided novel understanding of the molecular complexity of breast cancer. However, clinical decisions still rely on assessment of three molecular markers: estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor 2 (HER2) [1]. Identifying these markers is essential for efficient targeted treatment of the different types of breast cancer.
ER is a receptor that is activated by estrogen. There are two types of ER: ERα and ERβ. Once activated, ERα and ERβ form dimers and translocate into the nucleus to facilitate the regulation of various genes. Approximately 80% of breast cancer patients display ER-positive breast cancer. ER-positive breast cancer typically responds much more readily to hormone-targeted therapy. It is treated with tamoxifen, which is a drug that blocks hormone receptors, causing the inhibition of estrogen production [2].
PR is a receptor that blocks transcription until activated by progesterone. Approximately 65% of breast cancer patients display PR positive breast cancer. While PR may not respond to endocrine therapy directly, their activation may have significant impacts on the ER signaling pathway, thus showing value in their determination of which tumors may be subject to PR reprogramming of ER [2].
HER2 is a breast cancer diagnostic marker that aids the therapeutic decisions in the treatment of breast cancer [3]. Approximately 10-30% of patients display HER2 positive breast cancer. HER2 gene amplification in breast cancer cells is linked to a more clinically aggressive response in patients and corresponds to a higher death rate [4]. HER2 positive breast cancer is normally treated with trastuzumab, a monoclonal antibody that induces the downregulation and internalization of HER2, as well as upregulates cell cycle inhibitors [4].
Previously, detection of ER, PR, and HER2 in breast cancer cells has been widely done using the Immunohistochemistry (IHC) and Fluorescence in situ hybridization (FISH) methods. However, immunostaining methods such as IHC are naturally prone to error as the results are semi-quantitative and subject to interobserver variability [5]. Thus, these methods often abuse of neglect patients. FISH assessments give much more sensitive and quantitative results, but are expensive and time-consuming, thus not appropriate for widespread use. Furthermore, FISH methods are also prone to ambiguous results as standard protocols for sample collection and storage have not been developed yet.
Clearly, detection of receptor DNA or cDNA by PCR meets the criteria of speed and high sensitivity, so it is used frequently for diagnostic purposes. Gel-based reverse transcription polymerase chain reaction (RT-PCR) assay was reported for the diagnosis of African swine fever (ASF) [6,7]. This assay was sensitive and specific for the fast and early diagnosis of the ASF. Therefore, RT-PCR could be an alternative method for the detection of ER, PR, and HER2, as it is an inexpensive test that yields sensitive and accurate results [8]. Furthermore, multiplex PCR allows for simultaneous amplification of multiple target sequences in a single tube using specific primer sets in combination. Thus, multiplex PCR favors higher throughput and automation compared to singleplex PCR, especially when testing repeated and numerous patient analyses of the same targets [9]. To overcome the problems of two-step real-time singleplex RT-PCR, a highly sensitive and specific gel-based multiplex RT-PCR assay was developed in this study by testing ER, PR, and HER2 negative and positive breast cancer cell lines.
The Goal of the Project
- Develop simple and specific gel-based multiplex RT-PCR for detection of ER/PR/HER2-positive breast cancer using breast cancer cell line.
- Find the optimized multiplex RT-PCR condition by designing the primer sequences and adjusting melting and annealing temperature.
Method and Experiments
Breast cancer cell lines
4 different known breast cancer cell lines were used in this study as templates for the HER2 detection, being the MCF7, MDAMB231, BT474, and MDAMB453. These cell lines all display different types of cell markers (ER, PR, HER2), being HER2-, triple negative, triple positive, and HER2+ respectively (Table 1).
Primer set design
Primers were developed for ER, PR, HER2 and PUM1 (Table 2). They were chosen as all different lengths so they could be differentiated during agarose gel electrophoresis, with ER - 244 bp, PR - 224, HER2 - 342, and PUM1 - 187. Their annealing temperatures were fairly similar, all ranging around 60°C.
RT-PCR reaction
RNA was extracted from breast cancer cell lines by RNA-spin™ Total RNA Extraction Kit (Intron) following the manufacturer's instructions. cDNA was synthesized from the extracted RNA using TOPscript™ Reverse Transcriptase (Enzynomics) following the manufacturer’s instructions. For the preparation for the PCR reaction, the different primers were tested individually first using 20 µL reaction containing 2µL forward/reverse primers (10 pmol), 2µL reaction buffer, 2µL dNTP, 1.125µL Taq polymerase(Bioneer), 0.5 µL cDNA, and RNase Free dH2O up to 20 µL. PCR was done with annealing temperature 60°C and an extension time of 40 seconds in 72°C for 40 cycles. For the multiplex reactions, nTaq-multi HOT (Enzynomics) was used with 20 µL reaction containing 2µL forward/reverse primers, 0.2µL nTaq- multiHOT polymerase, 0.5 µL cDNA, and RNase Free dH2O up to 20 µL.
Agarose gel electrophoresis
2-5% agarose gel and TBE buffer were used in this study. RedSafe™ Nucleic Acid Staining Solution (Intron), which is an alternative to the traditional ethidium bromide(EtBr), was used to stain nucleic acid. Amplified DNA was detected by direct examination of the gel in ultraviolet(UV) light.
Agarose Gel Quantification Analysis
LI-COR Image Studio software version 2.1.10 and gel images were saved as a work area. All gel intensity quantification analyses were performed on images saved in TIFF format.
All experiments in this study including running the agarose gel electrophoresis were performed in University of Suwon, Korea.
Results
With the molecular classifications of breast cancer, researchers focus on breast cancer cell lines to determine whether the molecular profiles observed in breast cancer patients are reflected in cell line models of the disease. Applications of transcriptional profiling to breast cancer cell lines using various platforms provided the characteristics of the cell lines by the expression of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2). Therefore, 4 cell lines were selected to observe ER, PR, and HER2 expressions with RT-PCR method (Table 1).
Table 1. Categorization and molecular information of 4 breast cancer cell lines, and the clinical features of tumors where they derive
We categorize these cell lines with clinical features. AC is ‘adenocarcinoma’, and IDC is ‘invasive ductal carcinoma’.
Previous studies indicated that PUM1 is a suitable reference gene for normalization of gene expression levels in breast cancer. We designed the primer sets for ER, PR, HER2, and PUM1 as all different lengths so they could be differentiated during agarose gel electrophoresis (Table 2).
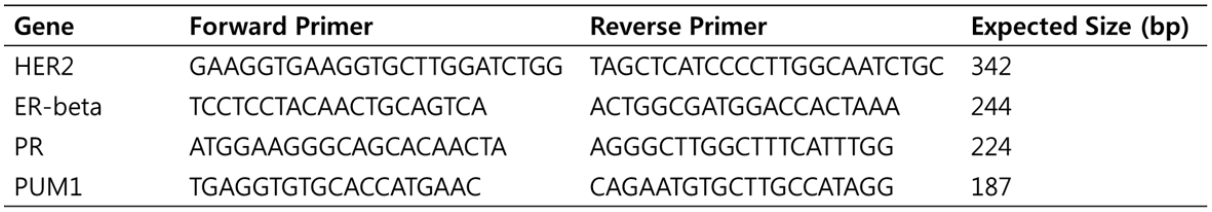
Table 2. Sequence information for primers used in RT-PCR
Previous studies indicated that PUM1 is a suitable reference gene for normalization of gene expression levels in breast cancer. We designed the primer sets for ER, PR, HER2, and PUM1 as all different lengths so they could be differentiated during agarose gel electrophoresis (Table 2).
Table 2. Sequence information for primers used in RT-PCR
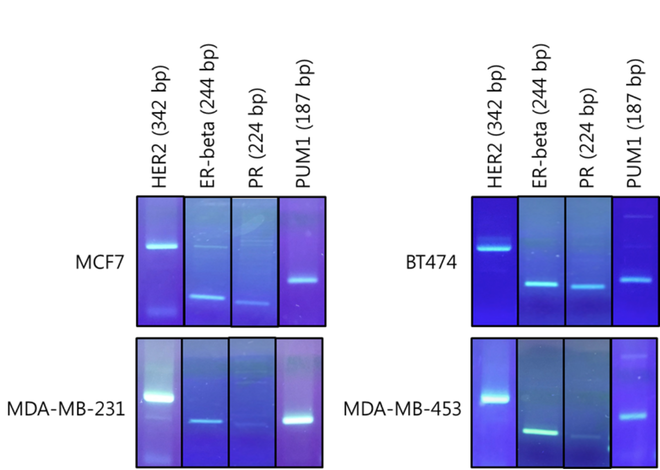
Figure 1. Agarose gel electrophoresis of singleplex RT-PCR products from cDNA of MCF7, BT-474, MDA-MB-231, and MDA-MB-453 breast cancer cell line.
The result of a PCR reaction is visualized using agarose gel electrophoresis. DNA fragments of the expected size of amplified HER2,ER, PR, and PUM1 form a single band on the gel by singleplex RT-PCR in all four breast cancer cell line (Figure 1). This result indicates that PCR conditions were optimized and the primers were highly specific and sensitive to amplify the targeted gene. The optimized RT-PCR reaction is specified in method and experiment section.
The result of a PCR reaction is visualized using agarose gel electrophoresis. DNA fragments of the expected size of amplified HER2,ER, PR, and PUM1 form a single band on the gel by singleplex RT-PCR in all four breast cancer cell line (Figure 1). This result indicates that PCR conditions were optimized and the primers were highly specific and sensitive to amplify the targeted gene. The optimized RT-PCR reaction is specified in method and experiment section.
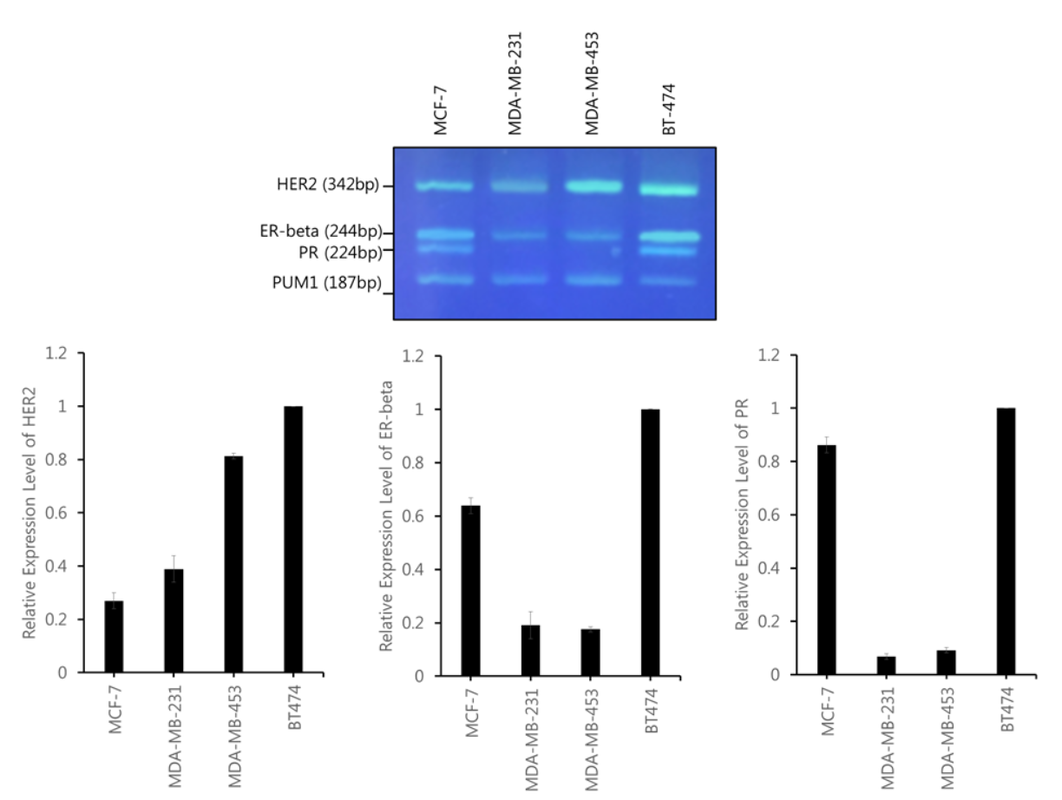
Figure 2. Quantification of HER2, ER-beta, and PR expression level on cDNA from MCF7, BT-474, MDA-MB-231, and MDA-MB-453 using agarose gel electrophoresis of multiplex RT-PCR. (upper) Gel electrophoresis of HER2 (342bp), ER-beta (244bp), PR (224bp), and PUM1 (187bp) (lower) Bar chart of band intensity calculated for HER2 expression level. The expression level of HER2, ER-beta, and PR was normalized by PUM1 expression level.
Since the PCR conditions were optimized in Figure 1, multiplex RT-PCR was performed in four different breast cancer cell lines. Multiplex RT-PCR showed viable results for all cell lines as the relative expression levels for ER, PR, and HER2 were low for the ER, PR, and HER2-negative breast cancer cell lines and high for the ER, PR, and HER2-positive breast cancer cell lines (Figure 2). PUM1 also proved to be very compatible with ER, PR, and HER2 in the multiplex process as the multiplex results yielded very clear and valid results. The observations show that gel-based RT-PCR multiplex detection of ER, PR, and HER2 is a reliable way of detecting the expression of ER, PR, and HER2 in breast cancer cell lines and have the potential to be further applied to other genes involved in breast cancer.
Since the PCR conditions were optimized in Figure 1, multiplex RT-PCR was performed in four different breast cancer cell lines. Multiplex RT-PCR showed viable results for all cell lines as the relative expression levels for ER, PR, and HER2 were low for the ER, PR, and HER2-negative breast cancer cell lines and high for the ER, PR, and HER2-positive breast cancer cell lines (Figure 2). PUM1 also proved to be very compatible with ER, PR, and HER2 in the multiplex process as the multiplex results yielded very clear and valid results. The observations show that gel-based RT-PCR multiplex detection of ER, PR, and HER2 is a reliable way of detecting the expression of ER, PR, and HER2 in breast cancer cell lines and have the potential to be further applied to other genes involved in breast cancer.
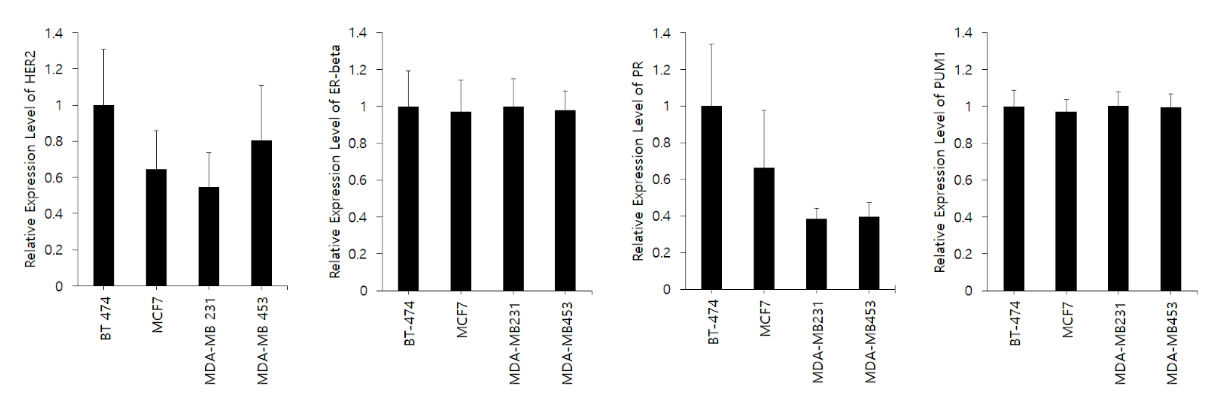
Figure 3. Relative expression for HER2, ER-beta, PR, and PUM1 from MCF7, BT-474, MDA-MB-231, and MDA-MB-453 using cell line microarray data provided by CellExpress.
CellExpress is a web-based tool that allows for analysis of gene expression levels in all of the cancer cell lines and clinical samples available online. The program takes queries based on gene, cell line, and normalization method and outputs all relevant data with a value corresponding to the relative expression of the gene and microRNA in the cell line in question. This system is highly useful for cross analysis of gene expression in different cell lines, as it provides a universal ranking and value with the same normalization method.
Figure 3 shows the relative expression of the four genes in this study in the MCF7, BT-474, MDA-MB-231, and MDA-MB-453 cell lines. PUM1 has a very uniform distribution of relative expression and has low standard deviations, suggesting it is an appropriate normalizing gene for this study. All microarray data is consistent with Figure 2 with the exception of the relative expression of ER-beta. The microarray data shows a much more even distribution of expression of ER-beta compared with Figure 2, which shows distinct differences in expression between the four cell lines. This could be explained by the fact that ER-beta has nine exons in total, and the selected region for the microarray and the primer used in this study could have targeted different exons in the gene. Thus, as there are many different isoforms of ER-beta, the relative expression could differ depending on the exon targeted.
Discussion
In triple negative breast cancer, there is a significantly lower patient survival rate because receptor-targeted hormonal therapy is unavailable to patients without sufficient expression of HER2, ER, and PR. Also, triple negative breast cancer is more likely to metastasize, which furthermore emphasizes the need for fast detection of receptor positive/negative cancer. In order to visualize the effects of triple negative breast cancer on patients, 14 studies and 9134 samples were compiled to construct data (Supplementary Figure 1) using a website called cBioPortal to analyze the difference in survival rate between patients with gene deletion in HER2, ER, and PR and patients without the deletion [10]. There was a markedly lower rate of survival in patients with gene deletion with a p-value of 1.237e-3. A previous study on a Brazillian cohort confirms that TNBC displays a more aggressive behavior, has a more frequent recurrence, and has a worse survival rate [11]. Therefore, these patients have to receive different treatment (i.e. chemotherapy), which is why it is crucial to detect the expression of these receptors in the early stages of cancer.
FISH and IHC have been widely accepted as the method of detecting ER, PR, and HER2, a critical factor in the treatment of invasive breast cancer patients. However, these methods both have limitations and are not perfectly suitable for the detection of ER, PR, and HER2. RT-qPCR detection of ER, PR, and HER2 has many advantages over these two methods, including cost, timeliness, accuracy, and sensitivity. The results in this study show that the gel-based RT-PCR assessments of ER, PR, and HER2 detection were generally very accurate in detecting the amplification of ER, PR, and HER2 in different known breast cancer cell lines (Figure 2). The results from this study are also consistent with various microarray data (Figure 3), further confirming the accuracy of this method. Therefore, ER, PR, and HER2 detection via gel-based RT-PCR can account for the disadvantages of methods like IHC and FISH in the future to aid rapid and accurate detection of ER, PR, and HER2 in a large number of samples. Since ER, PR, and HER2 expression level was quantitatively measured by gel-based multiplex RT-PCR, measuring the expression level of novel genes that are related to breast cancer progression by gel-based multiplex can be developed in the future.
A limitation of this study is that the qRT-PCR was tested only on four breast cancer cell lines instead of tissue samples from actual patients. Cancer cell lines were initially derived from tumors and cultured in two dimensional conditions. Due to the advantage of cell culture, it has been widely used as models to study molecular markers of cancer. However, data obtained from the cancer cell lines could be different from the data obtained from actual patient tissue samples as tissue samples are often heterogeneous, containing not only tumor cells, but also various degrees of normal cells such as red blood cells. Previous study found a strong correlation of the expression levels of ER (r=0.85) and PR (r=0.9) between IHC and RT-qPCR methods in patient tissue samples [12]. This result suggests that qRT-PCR is a promising complementary method to IHC for determining hormone receptors and protein markers used in breast cancer diagnosis.
Development of a multiplex RT-PCR assay for four different genes required a very careful selection of primers and manipulation of PCR conditions because multiple primers could potentially dimerize or suppress other primers. The program GenomeCompiler was used to construct 19 sets of primers, which were chosen based on the location of the exon, melting temperature, and the length of PCR product. In each gene, there are many different isoforms that consist of different combinations of exons. Therefore, primers must be chosen from a specific region in the gene that is conserved in most isoforms of the gene to eliminate biases related to uneven representation of isoforms. Ensembl, a web-based genome browser, was employed to visualize the exon structure and sequence for the four genes and exon regions that were highly conserved were selected for the primers. Furthermore, we ensured that the melting temperature of all primers were similar, and differentiated the length of PCR products in order to ensure separation for distinct detection during gel electrophoresis. Primers were tested individually to ensure the PCR condition was optimized, and later in different combinations in the span of over 500 PCR reactions in order to maximize the quality of the multi-PCR products.
Conclusion
The result of this study would help for rapid detection of ER, PR, and HER2 especially for laboratories that cannot afford a real-time PCR machine. Unlike FISH and IHC assay, this assay is much simpler, more convenient and easier to undertake. The multiplex analysis greatly reduces time-consuming procedures and eliminates additional manipulations that are normally required for FISH and IHC assays.
Reference
1. Dai, X., et al., Breast Cancer Cell Line Classification and Its Relevance with Breast Tumor Subtyping. J Cancer, 2017. 8(16): p. 3131-3141.
2. Parise, C.A. and V. Caggiano, Breast Cancer Survival Defined by the ER/PR/HER2 Subtypes and a Surrogate Classification according to Tumor Grade and Immunohistochemical Biomarkers. J Cancer Epidemiol, 2014. 2014: p. 469251.
3. Zhang, J. and Y. Liu, HER2 over-expression and response to different chemotherapy regimens in breast cancer. J Zhejiang Univ Sci B, 2008. 9(1): p. 5-9.
4. Gajria, D. and S. Chandarlapaty, HER2-amplified breast cancer: mechanisms of trastuzumab resistance and novel targeted therapies. Expert Rev Anticancer Ther, 2011. 11(2): p. 263-75.
5. Mrozkowiak, A., et al., HER2 status in breast cancer determined by IHC and FISH: comparison of the results. Pol J Pathol, 2004. 55(4): p. 165-71.
6. Aguero, M., et al., A highly sensitive and specific gel-based multiplex RT-PCR assay for the simultaneous and differential diagnosis of African swine fever and Classical swine fever in clinical samples. Vet Res, 2004. 35(5): p. 551-63.
7. Liu, L., et al., A one-step, gel-based RT-PCR assay with comparable performance to real-time RT-PCR for detection of classical swine fever virus. J Virol Methods, 2007. 139(2): p. 203-7.
8. El Hadi, H., et al., Development and evaluation of a novel RT-qPCR based test for the quantification of HER2 gene expression in breast cancer. Gene, 2017. 605: p. 114-122.
9. Hockman, D., et al., Comparison of multiplex PCR hybridization-based and singleplex real-time PCR-based assays for detection of low prevalence pathogens in spiked samples. J Microbiol Methods, 2017. 132: p. 76-82.
10. Gao, J., et al., Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal, 2013. 6(269): p. pl1.
11. Goncalves, H., Jr., et al., Survival Study of Triple-Negative and Non-Triple-Negative Breast Cancer in a Brazilian Cohort. Clin Med Insights Oncol, 2018. 12: p. 1179554918790563.
12. Christopherson, C., et al., Comparison of immunohistochemistry (IHC) and quantitative RT-PCR: ER, PR, and HER2 receptor status. Journal of Clinical Oncology, 2012. 30(27_suppl): p. 47-47.
CellExpress is a web-based tool that allows for analysis of gene expression levels in all of the cancer cell lines and clinical samples available online. The program takes queries based on gene, cell line, and normalization method and outputs all relevant data with a value corresponding to the relative expression of the gene and microRNA in the cell line in question. This system is highly useful for cross analysis of gene expression in different cell lines, as it provides a universal ranking and value with the same normalization method.
Figure 3 shows the relative expression of the four genes in this study in the MCF7, BT-474, MDA-MB-231, and MDA-MB-453 cell lines. PUM1 has a very uniform distribution of relative expression and has low standard deviations, suggesting it is an appropriate normalizing gene for this study. All microarray data is consistent with Figure 2 with the exception of the relative expression of ER-beta. The microarray data shows a much more even distribution of expression of ER-beta compared with Figure 2, which shows distinct differences in expression between the four cell lines. This could be explained by the fact that ER-beta has nine exons in total, and the selected region for the microarray and the primer used in this study could have targeted different exons in the gene. Thus, as there are many different isoforms of ER-beta, the relative expression could differ depending on the exon targeted.
Discussion
In triple negative breast cancer, there is a significantly lower patient survival rate because receptor-targeted hormonal therapy is unavailable to patients without sufficient expression of HER2, ER, and PR. Also, triple negative breast cancer is more likely to metastasize, which furthermore emphasizes the need for fast detection of receptor positive/negative cancer. In order to visualize the effects of triple negative breast cancer on patients, 14 studies and 9134 samples were compiled to construct data (Supplementary Figure 1) using a website called cBioPortal to analyze the difference in survival rate between patients with gene deletion in HER2, ER, and PR and patients without the deletion [10]. There was a markedly lower rate of survival in patients with gene deletion with a p-value of 1.237e-3. A previous study on a Brazillian cohort confirms that TNBC displays a more aggressive behavior, has a more frequent recurrence, and has a worse survival rate [11]. Therefore, these patients have to receive different treatment (i.e. chemotherapy), which is why it is crucial to detect the expression of these receptors in the early stages of cancer.
FISH and IHC have been widely accepted as the method of detecting ER, PR, and HER2, a critical factor in the treatment of invasive breast cancer patients. However, these methods both have limitations and are not perfectly suitable for the detection of ER, PR, and HER2. RT-qPCR detection of ER, PR, and HER2 has many advantages over these two methods, including cost, timeliness, accuracy, and sensitivity. The results in this study show that the gel-based RT-PCR assessments of ER, PR, and HER2 detection were generally very accurate in detecting the amplification of ER, PR, and HER2 in different known breast cancer cell lines (Figure 2). The results from this study are also consistent with various microarray data (Figure 3), further confirming the accuracy of this method. Therefore, ER, PR, and HER2 detection via gel-based RT-PCR can account for the disadvantages of methods like IHC and FISH in the future to aid rapid and accurate detection of ER, PR, and HER2 in a large number of samples. Since ER, PR, and HER2 expression level was quantitatively measured by gel-based multiplex RT-PCR, measuring the expression level of novel genes that are related to breast cancer progression by gel-based multiplex can be developed in the future.
A limitation of this study is that the qRT-PCR was tested only on four breast cancer cell lines instead of tissue samples from actual patients. Cancer cell lines were initially derived from tumors and cultured in two dimensional conditions. Due to the advantage of cell culture, it has been widely used as models to study molecular markers of cancer. However, data obtained from the cancer cell lines could be different from the data obtained from actual patient tissue samples as tissue samples are often heterogeneous, containing not only tumor cells, but also various degrees of normal cells such as red blood cells. Previous study found a strong correlation of the expression levels of ER (r=0.85) and PR (r=0.9) between IHC and RT-qPCR methods in patient tissue samples [12]. This result suggests that qRT-PCR is a promising complementary method to IHC for determining hormone receptors and protein markers used in breast cancer diagnosis.
Development of a multiplex RT-PCR assay for four different genes required a very careful selection of primers and manipulation of PCR conditions because multiple primers could potentially dimerize or suppress other primers. The program GenomeCompiler was used to construct 19 sets of primers, which were chosen based on the location of the exon, melting temperature, and the length of PCR product. In each gene, there are many different isoforms that consist of different combinations of exons. Therefore, primers must be chosen from a specific region in the gene that is conserved in most isoforms of the gene to eliminate biases related to uneven representation of isoforms. Ensembl, a web-based genome browser, was employed to visualize the exon structure and sequence for the four genes and exon regions that were highly conserved were selected for the primers. Furthermore, we ensured that the melting temperature of all primers were similar, and differentiated the length of PCR products in order to ensure separation for distinct detection during gel electrophoresis. Primers were tested individually to ensure the PCR condition was optimized, and later in different combinations in the span of over 500 PCR reactions in order to maximize the quality of the multi-PCR products.
Conclusion
The result of this study would help for rapid detection of ER, PR, and HER2 especially for laboratories that cannot afford a real-time PCR machine. Unlike FISH and IHC assay, this assay is much simpler, more convenient and easier to undertake. The multiplex analysis greatly reduces time-consuming procedures and eliminates additional manipulations that are normally required for FISH and IHC assays.
Reference
1. Dai, X., et al., Breast Cancer Cell Line Classification and Its Relevance with Breast Tumor Subtyping. J Cancer, 2017. 8(16): p. 3131-3141.
2. Parise, C.A. and V. Caggiano, Breast Cancer Survival Defined by the ER/PR/HER2 Subtypes and a Surrogate Classification according to Tumor Grade and Immunohistochemical Biomarkers. J Cancer Epidemiol, 2014. 2014: p. 469251.
3. Zhang, J. and Y. Liu, HER2 over-expression and response to different chemotherapy regimens in breast cancer. J Zhejiang Univ Sci B, 2008. 9(1): p. 5-9.
4. Gajria, D. and S. Chandarlapaty, HER2-amplified breast cancer: mechanisms of trastuzumab resistance and novel targeted therapies. Expert Rev Anticancer Ther, 2011. 11(2): p. 263-75.
5. Mrozkowiak, A., et al., HER2 status in breast cancer determined by IHC and FISH: comparison of the results. Pol J Pathol, 2004. 55(4): p. 165-71.
6. Aguero, M., et al., A highly sensitive and specific gel-based multiplex RT-PCR assay for the simultaneous and differential diagnosis of African swine fever and Classical swine fever in clinical samples. Vet Res, 2004. 35(5): p. 551-63.
7. Liu, L., et al., A one-step, gel-based RT-PCR assay with comparable performance to real-time RT-PCR for detection of classical swine fever virus. J Virol Methods, 2007. 139(2): p. 203-7.
8. El Hadi, H., et al., Development and evaluation of a novel RT-qPCR based test for the quantification of HER2 gene expression in breast cancer. Gene, 2017. 605: p. 114-122.
9. Hockman, D., et al., Comparison of multiplex PCR hybridization-based and singleplex real-time PCR-based assays for detection of low prevalence pathogens in spiked samples. J Microbiol Methods, 2017. 132: p. 76-82.
10. Gao, J., et al., Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal, 2013. 6(269): p. pl1.
11. Goncalves, H., Jr., et al., Survival Study of Triple-Negative and Non-Triple-Negative Breast Cancer in a Brazilian Cohort. Clin Med Insights Oncol, 2018. 12: p. 1179554918790563.
12. Christopherson, C., et al., Comparison of immunohistochemistry (IHC) and quantitative RT-PCR: ER, PR, and HER2 receptor status. Journal of Clinical Oncology, 2012. 30(27_suppl): p. 47-47.